Real-Time Illustration of Vascular StructuresFelix Ritter2006
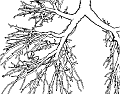
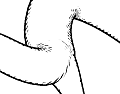
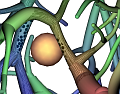
Understanding the branching pattern and topology of vascular structures is crucial for therapy planning and the actual surgery in order to prevent healthy organs or organ regions from being cut off from blood supply and drainage. A 3D visualization that provides knowledge about the location, properties, spatial distances, and functional relationships of those vessels to other relevant anatomic structures has been a frequent request by surgeons. While current therapy planning software can provide most of this information, an integrated visualization that enables the surgeon to make reliable judgments without time-consuming, interactive inspections still remains an open request. During surgery, the surgeon has even less time to analyze complex visualizations than at the planning stage. Ideally, such visualization would therefore be static in a sense of facilitating frequent look-ups of required information yet providing all necessary morphological and spatial information in one single picture. Such a picture could be printed, displayed on a monitor inside the operation theater, and eventually projected on the very organ before dissection. The perception of spatial distances, however, becomes demanding when viewing a static, monoscopic projection of a 3D visualization. This is especially true for complex vascular systems that may consist of multiple interweaved tree-like structures such as the vascular systems of the liver (portal vein, liver artery, hepatic veins, and biliary duct). The effectiveness and lucidity of the visualization highly depend on the accentuation of spatial depth as well as the perceptive separation of important, individual properties. To improve the communication of both aspects, the real-time vascular visualization methods presented in this paper utilize and extend on illustrative rendering techniques that provide functional realism. Illustrative visualization methods not only allow us to emphasize or omit properties, but also offer visualization techniques with limited use of color. Due to varying absorption and reflection characteristics on organ surfaces, the perceived color and brightness gradations resulting from a traditional shaded projection on the organ are difficult to predict, thus making them less suited for this purpose. Instead we propose the use of texture as an alternative visual attribute. This allows us to encode additional information, such as the local distance to a tumor.
|